Welcome
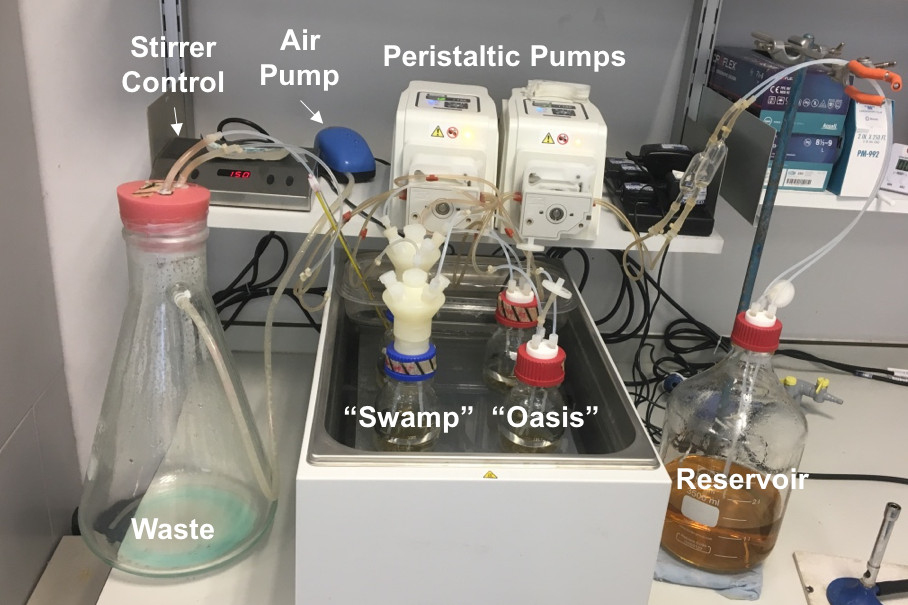
We are an experimental evolution lab. We use bacteriophages (esp. ϕX174) as model organisms to study evolutionary dynamics and the influence of ecological variables on these. Please find below a brief description of our lab's approaches and activities. Students, please look at the teaching menu for local and external resources.